H12/AOM/2/4¶
This page describes a signal of selection found in the
Angola An. coluzzii populationusing the H12 (Garud et al. 20XX) statistic.The focus of this signal is on chromosome arm
2R between positions 58,660,000 and
59,200,000.
The evidence supporting this signal is
strong ( >= 100 on both flanks).
>= 100 on both flanks).
Signal location. Blue markers show the values of the selection statistic. The dashed black line shows the fitted peak model. The shaded red area shows the focus of the selection signal. The shaded blue area shows the genomic region in linkage with the selection event. Use the mouse wheel or the controls at the top right of the plot to zoom in, and hover over genes to see gene names and descriptions.
Genes¶
The following 20 genes overlap the focal region: AGAP004630 (Erg28-domain containing protein), AGAP004631, AGAP004632 (DEF2 - defensin anti-microbial peptide), AGAP004633 (sodium-independent sulfate anion transporter), AGAP004634, AGAP004635 (sodium-independent sulfate anion transporter), AGAP013218 (sodium-independent sulfate anion transporter), AGAP004636 (sodium-independent sulfate anion transporter), AGAP004637 (transcriptional activator cubitus interruptus), AGAP013117, AGAP004638, AGAP004639, AGAP004640, AGAP013302, AGAP004641 (ATP synthase mitochondrial F1 complex assembly factor 1), AGAP004642 (NPF - Neuropeptide F), AGAP013063 (Synaptic vesicle protein), AGAP004643 (SCRB6 - Class B Scavenger Receptor (CD36 domain).), AGAP004644, AGAP004645 (juvenile hormone diol kinase).
The following 12 genes are within 50 kbp of the focal region: AGAP004619, AGAP004620 (Envelysin), AGAP004621, AGAP0046221 (glutamate dehydrogenase (NAD(P) )), AGAP004623 (anaphase-promoting complex subunit 6), AGAP004624, AGAP004625 (cortactin), AGAP004626, AGAP004627, AGAP004628 (scaffold protein salvador), AGAP004629, AGAP004646 (homeobox protein HoxA/B/C/D4).
Key to insecticide resistance candidate gene types: 1 metabolic; 2 target-site; 3 behavioural; 4 cuticular.
Diagnostics¶
The information below provides some diagnostics from the Peak modelling algorithm.
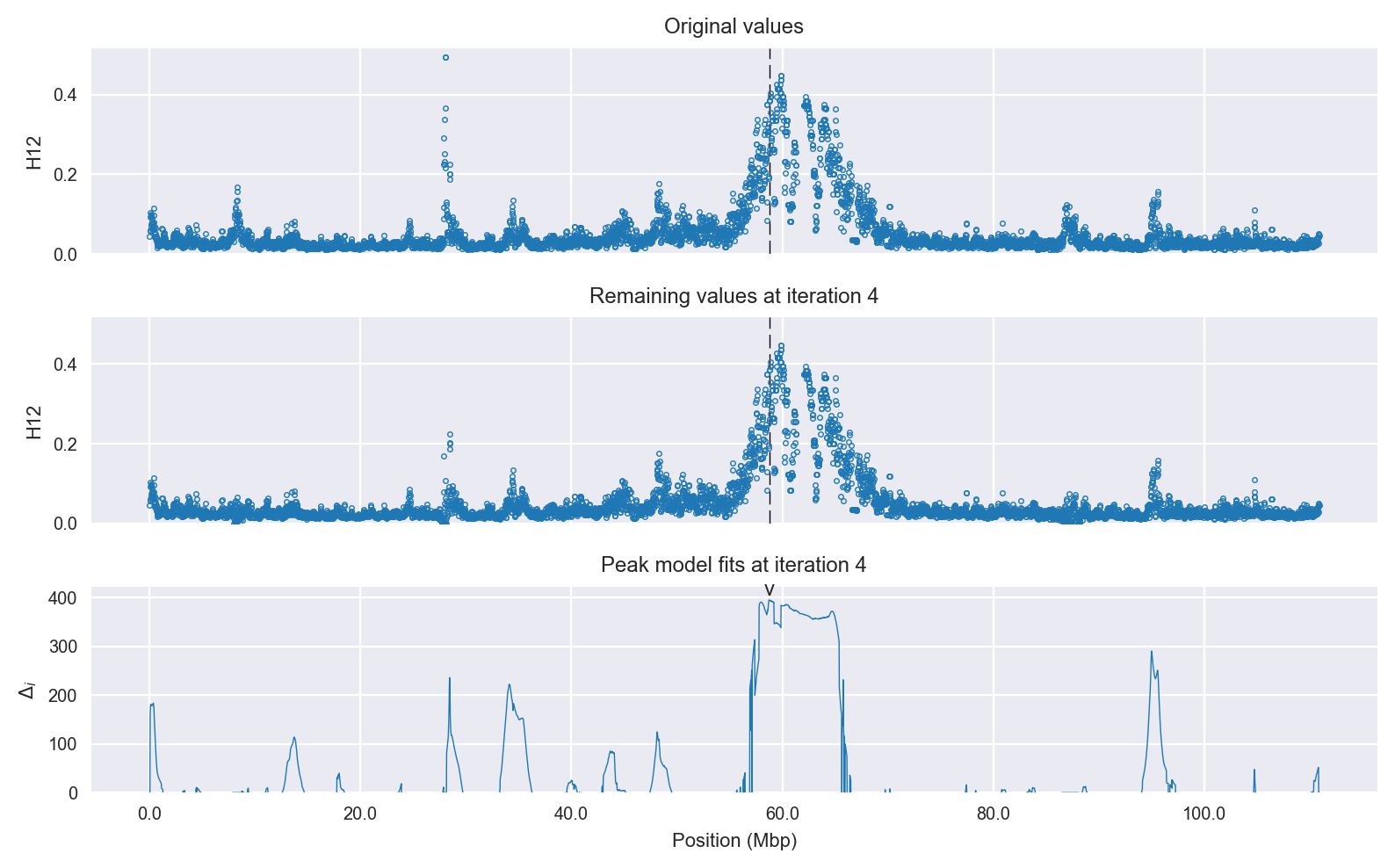
Selection signal in context. @@TODO
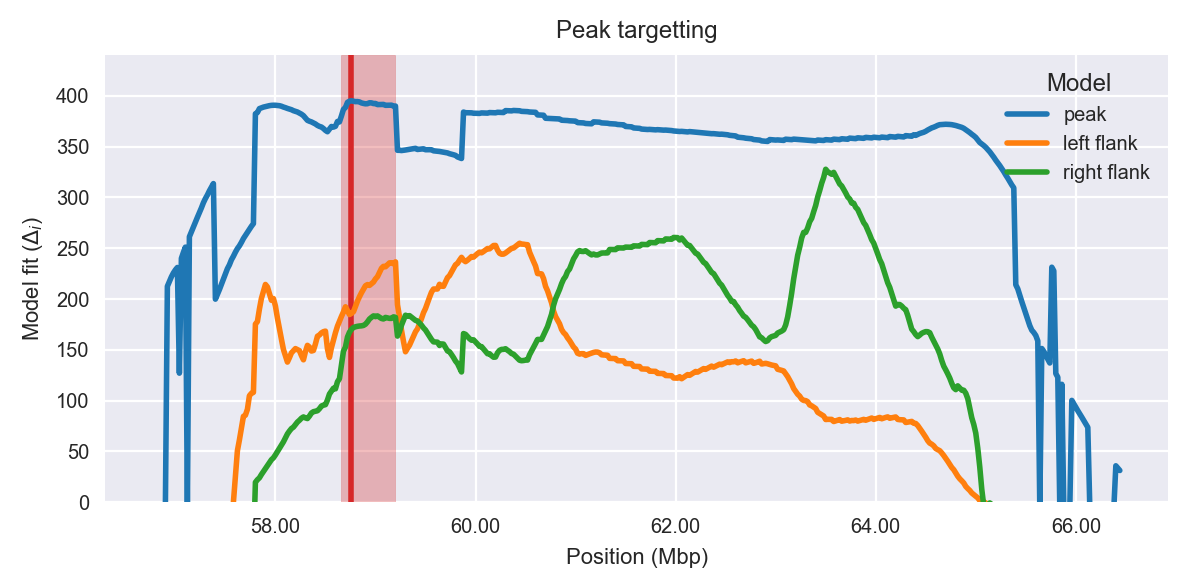
Peak targetting. @@TODO
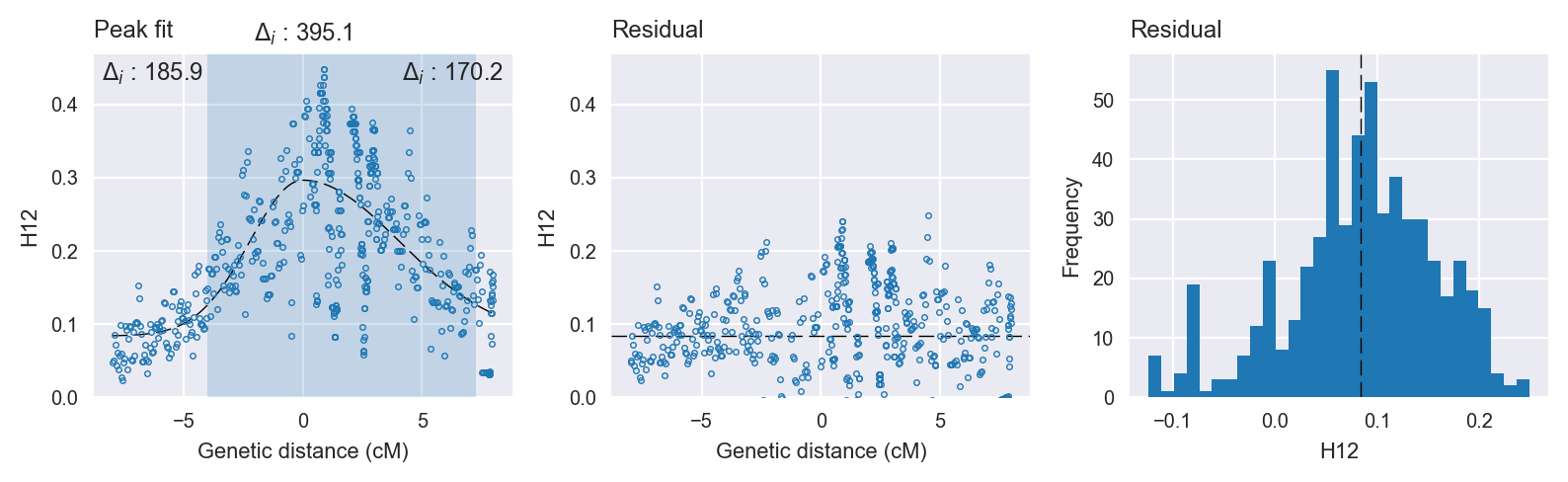
Peak fitting diagnostics. @@TODO
Model fit reports¶
Peak model:
[[Model]]
Model(skewed_gaussian)
[[Fit Statistics]]
# function evals = 72
# data points = 564
# variables = 4
chi-square = 3.051
reduced chi-square = 0.005
Akaike info crit = -2935.818
Bayesian info crit = -2918.478
[[Variables]]
center: 0 (fixed)
amplitude: 0.21263410 +/- 0.010066 (4.73%) (init= 0.5)
sigma: 2.99999788 +/- 0.201409 (6.71%) (init= 0.5)
skew: -0.43426841 +/- 0.052185 (12.02%) (init= 0)
baseline: 0.08373883 +/- 0.009670 (11.55%) (init= 0.03)
ceiling: 1 (fixed)
floor: 0 (fixed)
[[Correlations]] (unreported correlations are < 0.100)
C(amplitude, baseline) = -0.831
C(sigma, baseline) = -0.785
C(amplitude, sigma) = 0.472
C(sigma, skew) = 0.257
C(amplitude, skew) = 0.209
C(skew, baseline) = -0.171
Null model:
[[Model]]
Model(constant)
[[Fit Statistics]]
# function evals = 12
# data points = 563
# variables = 1
chi-square = 6.152
reduced chi-square = 0.011
Akaike info crit = -2540.760
Bayesian info crit = -2536.427
[[Variables]]
c: 0.20629218 +/- 0.004409 (2.14%) (init= 0.03)