H12/GWA/2/5¶
This page describes a signal of selection found in the
Guinea Bissau populationusing the H12 (Garud et al. 20XX) statistic.The focus of this signal is on chromosome arm
2R between positions 7,900,000 and
7,940,000.
The evidence supporting this signal is
weak ( < 50 on one or both flanks).
< 50 on one or both flanks).
Signal location. Blue markers show the values of the selection statistic. The dashed black line shows the fitted peak model. The shaded red area shows the focus of the selection signal. The shaded blue area shows the genomic region in linkage with the selection event. Use the mouse wheel or the controls at the top right of the plot to zoom in, and hover over genes to see gene names and descriptions.
Genes¶
Gene AGAP001674 (sidestep) overlaps the focal region.
Gene AGAP001675 is within 50 kbp of the focal region.
Key to insecticide resistance candidate gene types: 1 metabolic; 2 target-site; 3 behavioural; 4 cuticular.
Diagnostics¶
The information below provides some diagnostics from the Peak modelling algorithm.
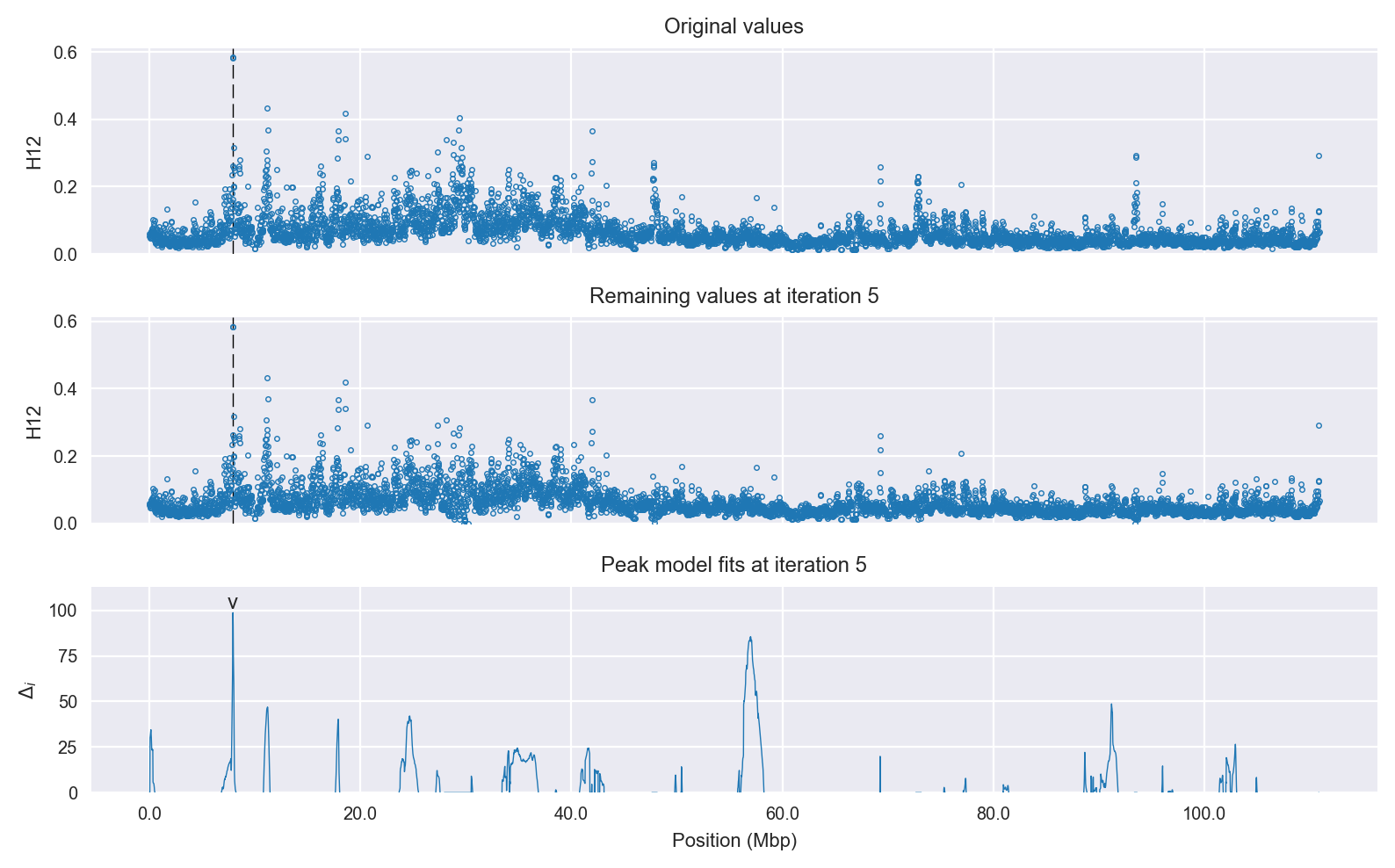
Selection signal in context. @@TODO
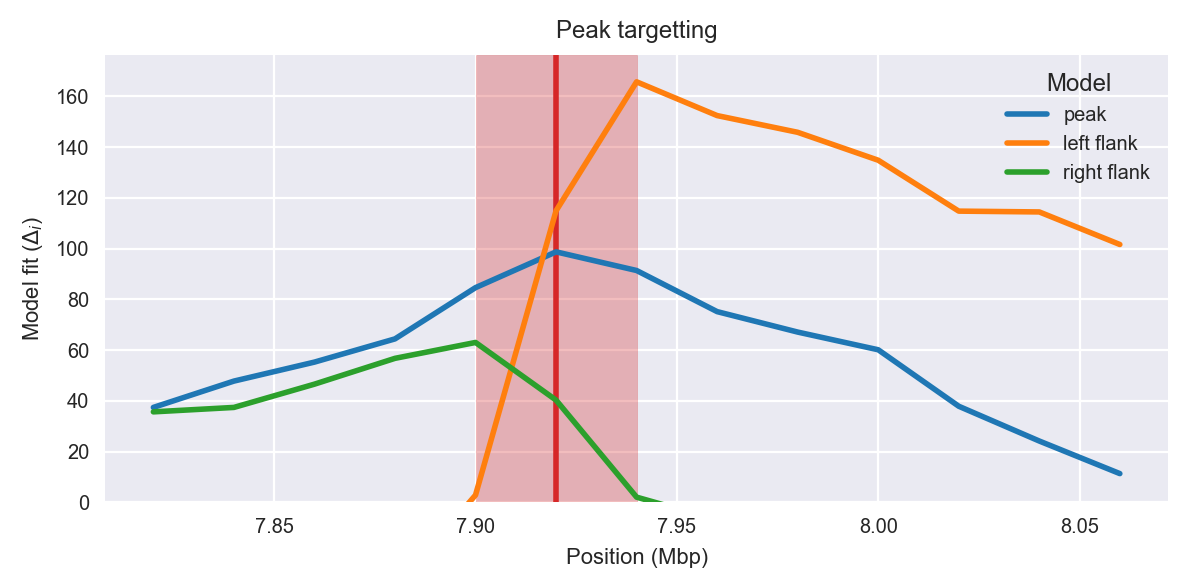
Peak targetting. @@TODO
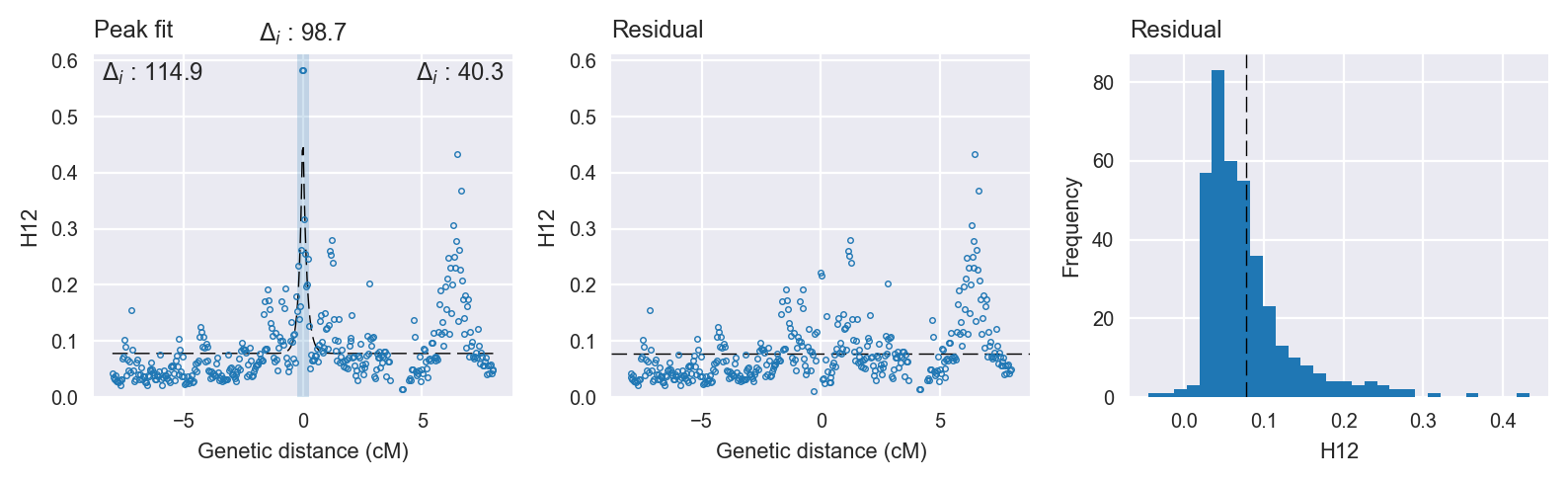
Peak fitting diagnostics. @@TODO
Model fit reports¶
Peak model:
[[Model]]
Model(skewed_exponential_peak)
[[Fit Statistics]]
# function evals = 44
# data points = 383
# variables = 4
chi-square = 1.259
reduced chi-square = 0.003
Akaike info crit = -2181.750
Bayesian info crit = -2165.958
[[Variables]]
center: 0 (fixed)
amplitude: 0.41560147 +/- 0.043080 (10.37%) (init= 0.5)
decay: 0.15000000 +/- 0.000108 (0.07%) (init= 0.5)
skew: -0.07725369 +/- 0.146211 (189.26%) (init= 0)
baseline: 0.07803526 +/- 0.003069 (3.93%) (init= 0.03)
ceiling: 1 (fixed)
floor: 0 (fixed)
[[Correlations]] (unreported correlations are < 0.100)
C(amplitude, decay) = -0.705
C(decay, baseline) = -0.203
Null model:
[[Model]]
Model(constant)
[[Fit Statistics]]
# function evals = 11
# data points = 382
# variables = 1
chi-square = 1.628
reduced chi-square = 0.004
Akaike info crit = -2083.052
Bayesian info crit = -2079.106
[[Variables]]
c: 0.08486079 +/- 0.003344 (3.94%) (init= 0.03)